About
1 Variant prediction
1.1 Input Instructions
PDB file can be downloaded by providing ID. Also the chain ID has to be provided. (e.g. >1aj3-A)
Provide the variants in HGVS format, i.e. original amino acid, position number, variant amino acid (e.g. P61A).
For prediction type, users can choose between regression and classification prediction.
1.2 Output
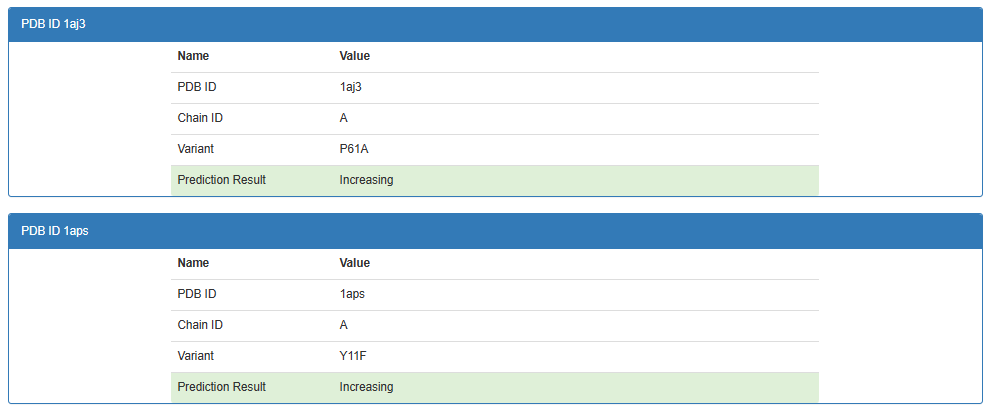
2 Protein prediction
Predict all possible 19 single amino acid substitutions in all positions in a single protein. Provide PDB and chain id.
2.1 Input Instructions
PDB file can be downloaded by providing ID. Also the chain ID has to be provided. (e.g. >2jof-A)
For prediction type, users can choose between regression and classification prediction.
It will take a long time, please provide your email and be patient.
2.2 Output
Prediction result for all position of the protein. example
Data sets
Extensive data mining was performed for obtaining cases for training and testing.
download
Dataset
variations
decreasing
no effect
increasing
Training dataset
762
520
136
106
Blind Test dataset
190
133
39
18
The source paper of the data sets.
| PDB ID | Number of variants | PubMed ID | |
|---|---|---|---|
| 1AJ3 | 2 | 15504411 | |
| 1APS | 24 | 10542090 | |
| 1BNR | 19 | 15035606 | |
| 1FKB | 34 | 10438631 | |
| 1HMK | 17 | 15276846 | |
| 1IMQ | 32 | 12547210 | |
| 1N88 | 17 | 15136744 | |
| 1SCE | 56 | 10673431 | |
| 1TEN | 39 | 10704314 | |
| 1TIT | 26 | 11377196 | |
| 2ABD | 30 | 10360367 | |
| 2CI2 | 68 | 7490748 | |
| 2PTL | 68 | 10801362 | |
| 3GB1 | 31 | 10932252 | |
| 1CSP | 22 | 15147842 | |
| 1DIV-C | 24 | 17240985 | |
| 1DIV-N | 14 | 16246369 | |
| 1E0L | 45 | 16784750 | |
| 1FMK | 57 | 10542092 | |
| 1O6X | 18 | 9799641 | |
| 1RFA | 47 | 17137592 | |
| 1SHF | 34 | 11786916 | |
| 1SHG | 14 | 10542091 | |
| 1SS1 | 63 | 17628591 | |
| 1SSO | 23 | 11124040 | |
| 1ST7 | 18 | 15690348 | |
| 1UBQ | 27 | 15857839 | |
| 1W4E | 22 | 16406408 | |
| 1W4J | 22 | 18625240 | |
| 1YYJ | 39 | 12069590 | |
Contact
If you have any problems, please contact Chong Zhang (20205227090@stu.suda.edu.cn).
